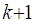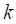Reading Data
The examples in this section illustrate various methods for
reading data into EPICURE.
Example
3.1 Reading Data from a CSV File with a Header Row and Creation of a BSF
File
In this example all of the
variables in the input file are to be read. Once the data have been read, the
SAVE command is used to write all of the variables to
a BSF file, which is given the default name TBFCHRT.BSF. Default BSF file names
are constructed by adding the BSF extension to the root of the input file name
(the root is that portion of the name that precedes the period [.], if any, in
the input file name). No transformations are carried out. The run is ended after
the save file is made. The complete set of commands necessary for this run is:
SESSION "ex3_1"
USETXT ../exdata/tbfchrt.csv @
INPUT @
SAVE @
The SESSION command instructs
EPICURE to write the session log and script (command) files called ex3_1.amf and
ex3_1.log, respectively. (The script file extension depends on the program
module in use when the session is started or when the SESSION command is given. For this example we used
AMFIT but we could have used any of the regression modules.) The USETXT
command reads the variable names from the header row of the CSV file. The INPUT
command tells the program to read the CSV file. (See Importing data from a CSV file
for an example of how to read these data using the menus.) The SAVE
command instructs the program to save the data in a BSF file. The file can be
saved from the menus or by using the toolbar
button  . If a BSF file with the
same name exists, you will be prompted to choose whether or not to overwrite the
file. The REPLACE subcommand of the SAVE command will force overwriting if a file with the same
name already exists. The output produced by the above commands is shown below.
. If a BSF file with the
same name exists, you will be prompted to choose whether or not to overwrite the
file. The REPLACE subcommand of the SAVE command will force overwriting if a file with the same
name already exists. The output produced by the above commands is shown below.
Output
3.2 Summary output from Example 3.1
USETXT ../exdata/tbfchrt.csv @
INPUT @
Input from ../exdata/tbfchrt.csv
1761 records
read 1760 records used
1 records rejected
20 variables defined At least 500 additional
variables can be created.
SAVE @
1760 records written to
../exdata/tbfchrt.BSF
Example 3.2 Reading Data from a Text File without Headers
and Creating a BSF file
If you wish to work with this new
example without restarted EPICURE, use menu or press the
Reset script toolbar button  . In this example all of the variables
in the input file are to be read. If, as in this file, the fields on each input
record are separated by one or more blanks and there is one record per
individual, there is no need to specify an input format. Once the data have been
read, the SAVE command is used to write all of the
variables to a BSF file. No transformations are carried out. The run is ended
after the file is saved. The complete set of commands necessary for this run is:
. In this example all of the variables
in the input file are to be read. If, as in this file, the fields on each input
record are separated by one or more blanks and there is one record per
individual, there is no need to specify an input format. Once the data have been
read, the SAVE command is used to write all of the
variables to a BSF file. No transformations are carried out. The run is ended
after the file is saved. The complete set of commands necessary for this run is:
NAMES id hosp byr bmo bdy inyr inmo indy tryr trmo
trdy
dose numbtf aft dxyr dxmo dxdy brstat
@
INPUT tbfchrt.dat @
SAVE @
END
The NAMES command provides names
for the input variables and indicates their position in the input file. The
INPUT command tells the program to read the data file.
The SAVE command instructs the program to save the
data in a BSF file. It can be executed from the menu items as described in the
previous example. Finally, the END command stops the program. The output
produced by the above commands is shown below.
Output 3.3 Summary output from Example 3.2
NAMES id hosp byr bmo bdy inyr inmo indy tryr trmo trdy
dose numbtf aft dxyr dxmo dxdy brstat @
INPUT ../exdata/tbfchrt.dat @
Input from ../exdata/tbfchrt.dat
1760 records
read 1760 records used
0 records rejected
20 variables defined At least 500 additional
variables can be created.
SAVE @
1760 records written to tbfchrt.BSF
Example
3.3 Importing data from a text file with a fixed format
The first step in reading data from a text file involves a
description of the input data. The data description commands in this example are
NAMES id hosp byr bmo bdy tryr trmo trdy
dose numbtf aft dxyr dxmo dxdy brstat
@
FORMAT
’(f6.0,1x,f1.0,f5.0,2f3.0,t29,3f3.0,f9.0,f5.0,
f6.0,3f3.0,f2.0)’@
The NAMES command is used here to
specify the names of the input variables, while the FORMAT command defines the
input format. In this example not all of the variables in the input file are
read (the study entry date variables are omitted), so we specify the input
format explicitly. Since the format string includes embedded blanks and is more
than one line long, it is enclosed in quotation marks.
NAMES and
FORMAT commands can be executed from the dialog window
that opens when we import data using the menu item (see also Importing data from other text
files ).

The format string resembles a FORTRAN format. For input
formats a field specification like F6.0 or I6 indicates that a number is to be read
from a six-column field. If the field specification is preceded by a
number, the specification will be repeated that many times. The 1X specifier indicates
that one column is to be skipped (the number before the x indicates the
number of columns to be skipped). The T29 specifier indicates that the next field
begins in column 29. Although not shown here, in FORTRAN formats the /
character is used to move to the next record and can be repeated to skip
records.
Format specifications can also be used when writing data to a
text file or to the screen using the DATA command. However, it is probably
best to specify variable-specific output formats using the SETFORMAT command or
to write the data as a CSV file using the NAMES and
COMMA options of the DATA
command. For output formats it is necessary to give the actual number of
digits to be written after the decimal point. For example F10.3 indicates
that the variable will be written in a field that is 10 columns wide with 3
digits after the decimal. There is a difference between Fx.0 and Ix output format
specifications. In both cases the number is written in a field that is
x characters
wide. However for the F format the number is rounded to the nearest
integer and the value is written with a trailing decimal point while for an I
format the value is truncated to an integer value and written without a decimal.
Note that EPICURE stores all data as double
precision floating point numbers.
Since the variables in this file
are separated by one or more blanks, it is possible to read these data without
an explicit format specification even though we want to skip some of the input
variables. This can be done using the command
NAMES id hosp byr bmo bdy „„ tryr trmo trdy
dose numbtf aft dxyr dxmo dxdy
brstat @
in place of the NAMES and FORMAT commands described above. The commas in this NAMES command indicate that three variables are to be
skipped. In general, 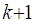 consecutive commas are used to
indicate that
consecutive commas are used to
indicate that 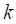 variables are to be
skipped.
variables are to be
skipped.
 . If a BSF file with the
same name exists, you will be prompted to choose whether or not to overwrite the
file. The REPLACE subcommand of the SAVE command will force overwriting if a file with the same
name already exists. The output produced by the above commands is shown below.
. If a BSF file with the
same name exists, you will be prompted to choose whether or not to overwrite the
file. The REPLACE subcommand of the SAVE command will force overwriting if a file with the same
name already exists. The output produced by the above commands is shown below.  . In this example all of the variables
in the input file are to be read. If, as in this file, the fields on each input
record are separated by one or more blanks and there is one record per
individual, there is no need to specify an input format. Once the data have been
read, the
. In this example all of the variables
in the input file are to be read. If, as in this file, the fields on each input
record are separated by one or more blanks and there is one record per
individual, there is no need to specify an input format. Once the data have been
read, the